Which vertices in graph can be reached in 1, 2 or N hops? This can basically be implemented in two ways. First, if the graph is implemented as adjacency matrix, by checking connections (ones in double array or a row) and iterating further.
Second, if the graph structure is implemented as ArrayList, by checking lists of adjacent vertices.
Both approaches result in long execution time, especially if the graph has many elements.
One alternative that I prefer is to do matrix multiplication on graph itself to find reachable vertices with N hops. Matrix multiplication performs much faster and the data within matrix data-structure is easier to manipulate.
Let’s suppose our matrix M has four vertices and some edges.
matrix([[0, 1, 1, 0],
[0, 0, 0, 1],
[0, 1, 0, 1],
[0, 0, 0, 0]])
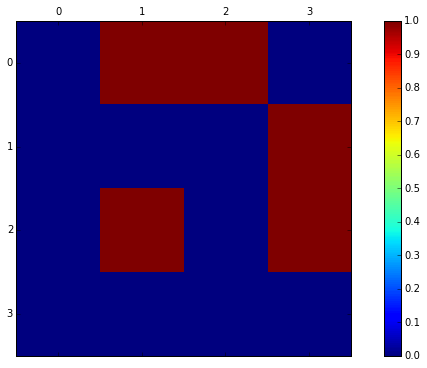
In order to find out which vertices are reachable within 2 hops from any given start-vertice we can simply multiply M by itself. M*M is shown below.
matrix([[0, 1, 0, 2],
[0, 0, 0, 0],
[0, 0, 0, 1],
[0, 0, 0, 0]])
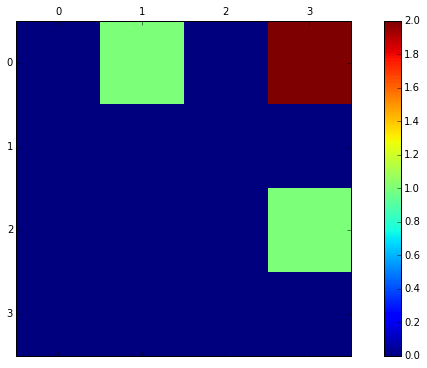
The Vertice 3(brown) is reachable from vertice 0 by 2 different paths in two hops. Vertices 1 and 3 (green) are reachable from vertices 0 and 2 within 2 hops by only one path.
If we wanted to check what vertices are reachable within 3 hops from any given start point, we simply do M^3.
matrix([[0, 0, 0, 1],
[0, 0, 0, 0],
[0, 0, 0, 0],
[0, 0, 0, 0]])
Only vertice 3 is reachable in 3 hops starting from vertice 0.
For N hops, M^N will do the trick.
Try it out with python. Below is the code to get started quickly.
## graph hops (Data Structures)
M = np.matrix([[0,1,1,0],
[0,0,0,1],
[0,1,0,1],
[0,0,0,0]])
def plot_matrix(matrix):
fig = plt.figure()
ax = fig.add_subplot(111)
cax = ax.matshow(matrix, interpolation='nearest')
fig.colorbar(cax)
plot_matrix(M)
plot_matrix(M^2)
plot_matrix(M^3)
Have fun and enjoy…

Leave a comment